تسلسل الجينوم الكامل
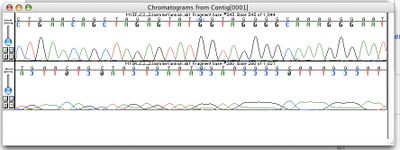

تسلسل الجينوم الكامل Whole genome sequencing، هو is ostensibly the process of determining the complete DNA sequence of an organism's genome at a single time. This entails sequencing all of an organism's chromosomal DNA as well as DNA contained in the mitochondria and, for plants, in the chloroplast. In practice, genome sequences that are nearly complete are also called whole genome sequences.[2]
Whole genome sequencing has largely been used as a research tool, but was being introduced to clinics in 2014.[3][4][5] In the future of personalized medicine, whole genome sequence data may be an important tool to guide therapeutic intervention.[6] The tool of gene sequencing at SNP level is also used to pinpoint functional variants from association studies and improve the knowledge available to researchers interested in evolutionary biology, and hence may lay the foundation for predicting disease susceptibility and drug response.
Whole genome sequencing should not be confused with DNA profiling, which only determines the likelihood that genetic material came from a particular individual or group, and does not contain additional information on genetic relationships, origin or susceptibility to specific diseases.[7] In addition, whole genome sequencing should not be confused with methods that sequence specific subsets of the genome - such methods include whole exome sequencing (1-2% of the genome) or SNP genotyping (<0.1% of the genome). As of 2017 there were no complete genomes for any mammals, including humans. Between 4% to 9% of the human genome, mostly satellite DNA, had not been sequenced.[8]
التاريخ





The DNA sequencing methods used in the 1970s and 1980s were manual; for example, Maxam–Gilbert sequencing and Sanger sequencing. Several whole bacteriophage and animal viral genomes were sequenced by these techniques, but the shift to more rapid, automated sequencing methods in the 1990s facilitated the sequencing of the larger bacterial and eukaryotic genomes.[10]
The first virus to have its complete genome sequenced was the Bacteriophage MS2 by 1976.[11] In 1992, yeast chromosome III was the first chromosome of any organism to be fully sequenced.[12] The first organism whose entire genome was fully sequenced was Haemophilus influenzae in 1995.[13] After it, the genomes of other bacteria and some archaea were first sequenced, largely due to their small genome size. H. influenzae has a genome of 1,830,140 base pairs of DNA.[13] In contrast, eukaryotes, both unicellular and multicellular such as Amoeba dubia and humans (Homo sapiens) respectively, have much larger genomes (see C-value paradox).[14] Amoeba dubia has a genome of 700 billion nucleotide pairs spread across thousands of chromosomes.[15] Humans contain fewer nucleotide pairs (about 3.2 billion in each germ cell – note the exact size of the human genome is still being revised) than A. dubia, however, their genome size far outweighs the genome size of individual bacteria.[16]
The first bacterial and archaeal genomes, including that of H. influenzae, were sequenced by Shotgun sequencing.[13] In 1996, the first eukaryotic genome (Saccharomyces cerevisiae) was sequenced. S. cerevisiae, a model organism in biology has a genome of only around 12 million nucleotide pairs,[17] and was the first unicellular eukaryote to have its whole genome sequenced. The first multicellular eukaryote, and animal, to have its whole genome sequenced was the nematode worm: Caenorhabditis elegans in 1998.[18] Eukaryotic genomes are sequenced by several methods including Shotgun sequencing of short DNA fragments and sequencing of larger DNA clones from DNA libraries such as bacterial artificial chromosomes (BACs) and yeast artificial chromosomes (YACs).[19]
In 1999, the entire DNA sequence of human chromosome 22, the second shortest human autosome, was published.[20] By the year 2000, the second animal and second invertebrate (yet first insect) genome was sequenced – that of the fruit fly Drosophila melanogaster – a popular choice of model organism in experimental research.[21] The first plant genome – that of the model organism Arabidopsis thaliana – was also fully sequenced by 2000.[22] By 2001, a draft of the entire human genome sequence was published.[23] The genome of the laboratory mouse Mus musculus was completed in 2002.[24]
In 2004, the Human Genome Project published an incomplete version of the human genome.[25] In 2008, a group from Leiden, the Netherlands, reported the sequencing of the first female human genome (Marjolein Kriek).
Currently thousands of genomes have been wholly or partially sequenced.
التفاصيل التجريبية
الخلايا المستخدمة للتسلسل
Almost any biological sample containing a full copy of the DNA—even a very small amount of DNA or ancient DNA—can provide the genetic material necessary for full genome sequencing. Such samples may include saliva, epithelial cells, bone marrow, hair (as long as the hair contains a hair follicle), seeds, plant leaves, or anything else that has DNA-containing cells.
The genome sequence of a single cell selected from a mixed population of cells can be determined using techniques of single cell genome sequencing. This has important advantages in environmental microbiology in cases where a single cell of a particular microorganism species can be isolated from a mixed population by microscopy on the basis of its morphological or other distinguishing characteristics. In such cases the normally necessary steps of isolation and growth of the organism in culture may be omitted, thus allowing the sequencing of a much greater spectrum of organism genomes.[26]
Single cell genome sequencing is being tested as a method of preimplantation genetic diagnosis, wherein a cell from the embryo created by in vitro fertilization is taken and analyzed before embryo transfer into the uterus.[27] After implantation, cell-free fetal DNA can be taken by simple venipuncture from the mother and used for whole genome sequencing of the fetus.[28]
التقنيات المبكرة

Sequencing of nearly an entire human genome was first accomplished in 2000 partly through the use of shotgun sequencing technology. While full genome shotgun sequencing for small (4000–7000 base pair) genomes was already in use in 1979,[29] broader application benefited from pairwise end sequencing, known colloquially as double-barrel shotgun sequencing. As sequencing projects began to take on longer and more complicated genomes, multiple groups began to realize that useful information could be obtained by sequencing both ends of a fragment of DNA. Although sequencing both ends of the same fragment and keeping track of the paired data was more cumbersome than sequencing a single end of two distinct fragments, the knowledge that the two sequences were oriented in opposite directions and were about the length of a fragment apart from each other was valuable in reconstructing the sequence of the original target fragment.
The first published description of the use of paired ends was in 1990 as part of the sequencing of the human HPRT locus,[30] although the use of paired ends was limited to closing gaps after the application of a traditional shotgun sequencing approach. The first theoretical description of a pure pairwise end sequencing strategy, assuming fragments of constant length, was in 1991.[31] In 1995, the innovation of using fragments of varying sizes was introduced,[32] and demonstrated that a pure pairwise end-sequencing strategy would be possible on large targets. The strategy was subsequently adopted by The Institute for Genomic Research (TIGR) to sequence the entire genome of the bacterium Haemophilus influenzae in 1995,[33] and then by Celera Genomics to sequence the entire fruit fly genome in 2000,[34] and subsequently the entire human genome. Applied Biosystems, now called Life Technologies, manufactured the automated capillary sequencers utilized by both Celera Genomics and The Human Genome Project.
التقنيات الحالية
While capillary sequencing was the first approach to successfully sequence a nearly full human genome, it is still too expensive and takes too long for commercial purposes. Since 2005 capillary sequencing has been progressively displaced by high-throughput (formerly "next-generation") sequencing technologies such as Illumina dye sequencing, pyrosequencing, and SMRT sequencing.[35] All of these technologies continue to employ the basic shotgun strategy, namely, parallelization and template generation via genome fragmentation.
Other technologies are emerging, including nanopore technology. Though nanopore sequencing technology is still being refined, its portability and potential capability of generating long reads are of relevance to whole-genome sequencing applications.[36]
التحليل
In principle, full genome sequencing can provide the raw nucleotide sequence of an individual organism's DNA. However, further analysis must be performed to provide the biological or medical meaning of this sequence, such as how this knowledge can be used to help prevent disease. Methods for analysing sequencing data are being developed and refined.
Because sequencing generates a lot of data (for example, there are approximately six billion base pairs in each human diploid genome), its output is stored electronically and requires a large amount of computing power and storage capacity.
While analysis of WGS data can be slow, it is possible to speed up this step by using dedicated hardware.[37]
تجارياً

A number of public and private companies are competing to develop a full genome sequencing platform that is commercially robust for both research and clinical use,[38] including Illumina,[39] Knome,[40] Sequenom,[41] 454 Life Sciences,[42] Pacific Biosciences,[43] Complete Genomics,[44] Helicos Biosciences,[45] GE Global Research (General Electric), Affymetrix, IBM, Intelligent Bio-Systems,[46] Life Technologies, Oxford Nanopore Technologies,[47] and the Beijing Genomics Institute.[48][49][50] These companies are heavily financed and backed by venture capitalists, hedge funds, and investment banks.[51][52]
A commonly-referenced commercial target for sequencing cost until the late 2010s was $1,000 USD, however, the private companies are working to reach a new target of only $100.[53]
الحافز
In October 2006, the X Prize Foundation, working in collaboration with the J. Craig Venter Science Foundation, established the Archon X Prize for Genomics,[54] intending to award $10 million to "the first team that can build a device and use it to sequence 100 human genomes within 10 days or less, with an accuracy of no more than one error in every 1,000,000 bases sequenced, with sequences accurately covering at least 98% of the genome, and at a recurring cost of no more than $1,000 per genome".[55] The Archon X Prize for Genomics was cancelled in 2013, before its official start date.[56][57]
التاريخ
In 2007, Applied Biosystems started selling a new type of sequencer called SOLiD System.[58] The technology allowed users to sequence 60 gigabases per run.[59]
In June 2009, Illumina announced that they were launching their own Personal Full Genome Sequencing Service at a depth of 30× for $48,000 per genome.[60][61] In August, the founder of Helicos Biosciences, Stephen Quake, stated that using the company's Single Molecule Sequencer he sequenced his own full genome for less than $50,000.[62] In November, Complete Genomics published a peer-reviewed paper in Science demonstrating its ability to sequence a complete human genome for $1,700.[63][64]
In May 2011, Illumina lowered its Full Genome Sequencing service to $5,000 per human genome, or $4,000 if ordering 50 or more.[65] Helicos Biosciences, Pacific Biosciences, Complete Genomics, Illumina, Sequenom, ION Torrent Systems, Halcyon Molecular, NABsys, IBM, and GE Global appear to all be going head to head in the race to commercialize full genome sequencing.[35][66]
With sequencing costs declining, a number of companies began claiming that their equipment would soon achieve the $1,000 genome: these companies included Life Technologies in January 2012,[67] Oxford Nanopore Technologies in February 2012,[68] and Illumina in February 2014.[69][70] In 2015, the NHGRI estimated the cost of obtaining a whole-genome sequence at around $1,500.[71] In 2016, Veritas Genetics began selling whole genome sequencing, including a report as to some of the information in the sequencing for $999.[72] In summer 2019, Veritas Genetics cut the cost for WGS to $599.[73] In 2017, BGI began offering WGS for $600.[74]
However, in 2015, some noted that effective use of whole gene sequencing can cost considerably more than $1000.[75] Also, reportedly there remain parts of the human genome that have not been fully sequenced by 2017.[76][77]
مقارنة بتقنيات أخرى
مصفوفات الدنا المكروية
Full genome sequencing provides information on a genome that is orders of magnitude larger than by DNA arrays, the previous leader in genotyping technology.
For humans, DNA arrays currently provide genotypic information on up to one million genetic variants,[78][79][80] while full genome sequencing will provide information on all six billion bases in the human genome, or 3,000 times more data. Because of this, full genome sequencing is considered a disruptive innovation to the DNA array markets as the accuracy of both range from 99.98% to 99.999% (in non-repetitive DNA regions) and their consumables cost of $5000 per 6 billion base pairs is competitive (for some applications) with DNA arrays ($500 per 1 million basepairs).[42]
التطبيقات
تسلسلات الطفرات
Whole genome sequencing has established the mutation frequency for whole human genomes. The mutation frequency in the whole genome between generations for humans (parent to child) is about 70 new mutations per generation.[81][82] An even lower level of variation was found comparing whole genome sequencing in blood cells for a pair of monozygotic (identical twins) 100-year-old centenarians.[83] Only 8 somatic differences were found, though somatic variation occurring in less than 20% of blood cells would be undetected.
In the specifically protein coding regions of the human genome, it is estimated that there are about 0.35 mutations that would change the protein sequence between parent/child generations (less than one mutated protein per generation).[84]
In cancer, mutation frequencies are much higher, due to genome instability. This frequency can further depend on patient age, exposure to DNA damaging agents (such as UV-irradiation or components of tobacco smoke) and the activity/inactivity of DNA repair mechanisms.[citation needed] Furthermore, mutation frequency can vary between cancer types: in germline cells, mutation rates occur at approximately 0.023 mutations per megabase, but this number is much higher in breast cancer (1.18-1.66 somatic mutations per Mb), in lung cancer (17.7) or in melanomas (≈33).[85] Since the haploid human genome consists of approximately 3,200 megabases,[86] this translates into about 74 mutations (mostly in noncoding regions) in germline DNA per generation, but 3,776-5,312 somatic mutations per haploid genome in breast cancer, 56,640 in lung cancer and 105,600 in melanomas.
The distribution of somatic mutations across the human genome is very uneven,[87] such that the gene-rich, early-replicating regions receive fewer mutations than gene-poor, late-replicating heterochromatin, likely due to differential DNA repair activity.[88] In particular, the histone modification H3K9me3 is associated with high,[89] and H3K36me3 with low mutation frequencies.[90]
الدرسات المرتبطة بالجينوم
In research, whole-genome sequencing can be used in a Genome-Wide Association Study (GWAS) - a project aiming to determine the genetic variant or variants associated with a disease or some other phenotype.[91]
الاستخدام التشخيصي
In 2009, Illumina released its first whole genome sequencers that were approved for clinical as opposed to research-only use and doctors at academic medical centers began quietly using them to try to diagnose what was wrong with people whom standard approaches had failed to help.[92] The price to sequence a genome at that time was US$19,500, which was billed to the patient but usually paid for out of a research grant; one person at that time had applied for reimbursement from their insurance company.[92] For example, one child had needed around 100 surgeries by the time he was three years old, and his doctor turned to whole genome sequencing to determine the problem; it took a team of around 30 people that included 12 bioinformatics experts, three sequencing technicians, five physicians, two genetic counsellors and two ethicists to identify a rare mutation in the XIAP that was causing widespread problems.[92][93][94]
Due to recent cost reductions (see above) whole genome sequencing has become a realistic application in DNA diagnostics. In 2013, the 3Gb-TEST consortium obtained funding from the European Union to prepare the health care system for these innovations in DNA diagnostics.[95][96] Quality assessment schemes, Health technology assessment and guidelines have to be in place. The 3Gb-TEST consortium has identified the analysis and interpretation of sequence data as the most complicated step in the diagnostic process.[97] At the Consortium meeting in Athens in September 2014, the Consortium coined the word genotranslation for this crucial step. This step leads to a so-called genoreport. Guidelines are needed to determine the required content of these reports.
Currently available newborn screening for childhood diseases allows detection of rare disorders that can be prevented or better treated by early detection and intervention. Specific genetic tests are also available to determine an etiology when a child's symptoms appear to have a genetic basis. Full genome sequencing, in addition has the potential to reveal a large amount of information (such as carrier status for autosomal recessive disorders, genetic risk factors for complex adult-onset diseases, and other predictive medical and non-medical information) that is currently not completely understood, may not be clinically useful to the child during childhood, and may not necessarily be wanted by the individual upon reaching adulthood.
This article needs additional citations for verification. (February 2017) (Learn how and when to remove this template message) |
Genomes2People (G2P), an initiative of Brigham and Women's Hospital and Harvard Medical School was created in 2011 to examine the integration of genomic sequencing into clinical care of adults and children.[98] G2P's director, Robert C. Green, had previously led the REVEAL study — Risk Evaluation and Education for Alzheimer's Disease – a series of clinical trials exploring patient reactions to the knowledge of their genetic risk for Alzheimer's.[99][100]
In 2018, researchers at Rady Children's Institute for Genomic Medicine in San Diego, CA determined that rapid whole-genome sequencing (rWGS) can diagnose genetic disorders in time to change acute medical or surgical management (clinical utility) and improve outcomes in acutely ill infants. The researchers reported a retrospective cohort study of acutely ill inpatient infants in a regional children's hospital from July 2016-March 2017. Forty-two families received rWGS for etiologic diagnosis of genetic disorders. The diagnostic sensitivity of rWGS was 43% (eighteen of 42 infants) and 10% (four of 42 infants) for standard genetic tests (P = .0005). The rate of clinical utility of rWGS (31%, thirteen of 42 infants) was significantly greater than for standard genetic tests (2%, one of 42; P = .0015). Eleven (26%) infants with diagnostic rWGS avoided morbidity, one had a 43% reduction in likelihood of mortality, and one started palliative care. In six of the eleven infants, the changes in management reduced inpatient cost by $800,000-$2,000,000. These findings replicate a prior study of the clinical utility of rWGS in acutely ill inpatient infants, and demonstrate improved outcomes and net healthcare savings. rWGS merits consideration as a first tier test in this setting.[101]
الاعتبارات الأخلاقية
Public human genome sequences
First people with public genome sequences
The first nearly complete human genomes sequenced were two Americans of predominantly Northwestern European ancestry in 2007 (J. Craig Venter at 7.5-fold coverage,[102][103][104] and James Watson at 7.4-fold).[105][106][107] This was followed in 2008 by sequencing of an anonymous Han Chinese man (at 36-fold),[108] a Yoruban man from Nigeria (at 30-fold),[109] a female clinical geneticist (Marjolein Kriek) from the Netherlands (at 7 to 8-fold), and a female leukemia patient in her mid-50s (at 33 and 14-fold coverage for tumor and normal tissues).[110] Steve Jobs was among the first 20 people to have their whole genome sequenced, reportedly for the cost of $100,000.[111] اعتبارا من يونيو 2012[تحديث], there were 69 nearly complete human genomes publicly available.[112] In November 2013, a Spanish family made their personal genomics data publicly available under a Creative Commons public domain license. The work was led by Manuel Corpas and the data obtained by direct-to-consumer genetic testing with 23andMe and the Beijing Genomics Institute. This is believed to be the first such Public Genomics dataset for a whole family.[113]
Databases
According to Science, the major databases of whole genomes are:[114]
| Biobank | Completed whole genomes | Release/access information |
|---|---|---|
| UK Biobank | 500,000 | Made available through a Web platform in November 2021, it is the largest public dataset of whole genomes. The genomes are linked to anonymized medical information and are made more accessible for biomedical research than prior, less comprehensive datasets. 300,000 more genomes were released in early 2023.[114][115][116] |
| Trans-Omics for Precision Medicine | 161,000 | National Institutes of Health (NIH) requires project-specific consent |
| Million Veteran Program | 125,000 | Non–Veterans Affairs researchers get access in 2022 |
| Genomics England's 100,000 Genomes | 120,000 | Researchers must join collaboration |
| All of Us | 90,000 | NIH expects to release by early 2022 |
انظر أيضاً
- Coverage (genetics)
- Whole Exome Sequencing
- DNA sequencing
- DNA microarray
- DNA profiling
- Horizontal correlation
- Medical genetics
- Nucleic acid sequence
- Human Genome Project
- Personal Genome Project
- Genomics England
- Predictive medicine
- Personalized medicine
- Duplex sequencing
جينوم عُرف تسلسلها
المصادر
- ^ Alberts, Bruce; Johnson, Alexander; Lewis, Julian; Raff, Martin; Roberts, Keith; Walter, Peter (2008). "Chapter 8". Molecular biology of the cell (5th ed.). New York: Garland Science. p. 550. ISBN 978-0-8153-4106-2.
- ^ "Definition of whole-genome sequencing - NCI Dictionary of Cancer Terms". National Cancer Institute (in الإنجليزية). 2012-07-20. Retrieved 2018-10-13.
- ^ Gilissen (Jul 2014). "Genome sequencing identifies major causes of severe intellectual disability". Nature. 511 (7509): 344–7. Bibcode:2014Natur.511..344G. doi:10.1038/nature13394. PMID 24896178.
- ^ Nones, K; Waddell, N; Wayte, N; Patch, AM; Bailey, P; Newell, F; Holmes, O; Fink, JL; Quinn, MC; Tang, YH; Lampe, G; Quek, K; Loffler, KA; Manning, S; Idrisoglu, S; Miller, D; Xu, Q; Waddell, N; Wilson, PJ; Bruxner, TJ; Christ, AN; Harliwong, I; Nourse, C; Nourbakhsh, E; Anderson, M; Kazakoff, S; Leonard, C; Wood, S; Simpson, PT; Reid, LE; Krause, L; Hussey, DJ; Watson, DI; Lord, RV; Nancarrow, D; Phillips, WA; Gotley, D; Smithers, BM; Whiteman, DC; Hayward, NK; Campbell, PJ; Pearson, JV; Grimmond, SM; Barbour, AP (29 October 2014). "Genomic catastrophes frequently arise in esophageal adenocarcinoma and drive tumorigenesis". Nature Communications. 5: 5224. Bibcode:2014NatCo...5.5224N. doi:10.1038/ncomms6224. PMC 4596003. PMID 25351503.
- ^ van El, CG; Cornel, MC; Borry, P; Hastings, RJ; Fellmann, F; Hodgson, SV; Howard, HC; Cambon-Thomsen, A; Knoppers, BM; Meijers-Heijboer, H; Scheffer, H; Tranebjaerg, L; Dondorp, W; de Wert, GM (June 2013). "Whole-genome sequencing in health care. Recommendations of the European Society of Human Genetics". European Journal of Human Genetics. 21 Suppl 1: S1–5. doi:10.1038/ejhg.2013.46. PMC 3660957. PMID 23819146.
- ^ Mooney, Sean (Sep 2014). "Progress towards the integration of pharmacogenomics in practice". Human Genetics. 134 (5): 459–65. doi:10.1007/s00439-014-1484-7. PMC 4362928. PMID 25238897.
- ^ Kijk magazine, 01 January 2009
- ^ "Psst, the human genome was never completely sequenced". STAT (in الإنجليزية الأمريكية). 2017-06-20. Archived from the original on 2017-10-23. Retrieved 2017-10-23.
- ^ Marx, Vivien (11 September 2013). "Next-generation sequencing: The genome jigsaw". Nature. 501 (7466): 263–268. Bibcode:2013Natur.501..263M. doi:10.1038/501261a. PMID 24025842.
- ^ al.], Bruce Alberts ... [et (2008). Molecular biology of the cell (5th ed.). New York: Garland Science. p. 551. ISBN 978-0-8153-4106-2.
- ^ Fiers, W.; Contreras, R.; Duerinck, F.; Haegeman, G.; Iserentant, D.; Merregaert, J.; Min Jou, W.; Molemans, F.; Raeymaekers, A.; Van den Berghe, A.; Volckaert, G.; Ysebaert, M. (8 April 1976). "Complete nucleotide sequence of bacteriophage MS2 RNA: primary and secondary structure of the replicase gene". Nature. 260 (5551): 500–507. Bibcode:1976Natur.260..500F. doi:10.1038/260500a0. PMID 1264203. S2CID 4289674.
- ^ Oliver, S. G.; van der Aart, Q. J. M.; Agostoni-Carbone, M. L.; et al. (May 1992). "The complete DNA sequence of yeast chromosome III". Nature. 357 (6373): 38–46. Bibcode:1992Natur.357...38O. doi:10.1038/357038a0. PMID 1574125. S2CID 4271784.
- ^ أ ب ت Fleischmann, R.; Adams, M.; White, O; Clayton, R.; Kirkness, E.; Kerlavage, A.; Bult, C.; Tomb, J.; Dougherty, B.; Merrick, J.; al., e. (28 July 1995). "Whole-genome random sequencing and assembly of Haemophilus influenzae Rd". Science. 269 (5223): 496–512. Bibcode:1995Sci...269..496F. doi:10.1126/science.7542800. PMID 7542800.
- ^ Eddy, Sean R. (November 2012). "The C-value paradox, junk DNA and ENCODE". Current Biology. 22 (21): R898–R899. Bibcode:2012CBio...22.R898E. doi:10.1016/j.cub.2012.10.002. PMID 23137679.
- ^ Pellicer, Jaume; FAY, Michael F.; Leitch, Ilia J. (15 September 2010). "The largest eukaryotic genome of them all?". Botanical Journal of the Linnean Society. 164 (1): 10–15. doi:10.1111/j.1095-8339.2010.01072.x.
- ^ Human Genome Sequencing Consortium, International (21 October 2004). "Finishing the euchromatic sequence of the human genome". Nature. 431 (7011): 931–945. Bibcode:2004Natur.431..931H. doi:10.1038/nature03001. PMID 15496913.
- ^ Goffeau, A.; Barrell, B. G.; Bussey, H.; Davis, R. W.; Dujon, B.; Feldmann, H.; Galibert, F.; Hoheisel, J. D.; Jacq, C.; Johnston, M.; Louis, E. J.; Mewes, H. W.; Murakami, Y.; Philippsen, P.; Tettelin, H.; Oliver, S. G. (25 October 1996). "Life with 6000 Genes". Science. 274 (5287): 546–567. Bibcode:1996Sci...274..546G. doi:10.1126/science.274.5287.546. PMID 8849441. S2CID 16763139. Archived (PDF) from the original on 7 March 2016.
- ^ The C. elegans Sequencing Consortium (11 December 1998). "Genome Sequence of the Nematode C. elegans: A Platform for Investigating Biology". Science. 282 (5396): 2012–2018. Bibcode:1998Sci...282.2012.. doi:10.1126/science.282.5396.2012. PMID 9851916.
- ^ Alberts, Bruce (2008). Molecular Biology of the Cell (5th ed.). New York: Garland Science. p. 552. ISBN 978-0-8153-4106-2.
- ^ Dunham, I. (December 1999). "The DNA sequence of human chromosome 22". Nature. 402 (6761): 489–495. Bibcode:1999Natur.402..489D. doi:10.1038/990031. PMID 10591208.
- ^ Adams MD; Celniker SE; Holt RA; et al. (2000-03-24). "The Genome Sequence of Drosophila melanogaster". Science. 287 (5461): 2185–2195. Bibcode:2000Sci...287.2185.. CiteSeerX 10.1.1.549.8639. doi:10.1126/science.287.5461.2185. PMID 10731132.
- ^ The Arabidopsis Genome Initiative (2000-12-14). "Analysis of the genome sequence of the flowering plant Arabidopsis thaliana". Nature. 408 (6814): 796–815. Bibcode:2000Natur.408..796T. doi:10.1038/35048692. PMID 11130711.
- ^ Venter JC; Adams MD; Myers EW; et al. (2001-02-16). "The Sequence of the Human Genome". Science. 291 (5507): 1304–1351. Bibcode:2001Sci...291.1304V. doi:10.1126/science.1058040. PMID 11181995.
- ^ Waterston RH; Lindblad-Toh K; Birney E; et al. (2002-10-31). "Initial sequencing and comparative analysis of the mouse genome". Nature. 420 (6915): 520–562. Bibcode:2002Natur.420..520W. doi:10.1038/nature01262. PMID 12466850.
- ^ International Human Genome Sequencing Consortium (2004-09-07). "Finishing the euchromatic sequence of the human genome". Nature. 431 (7011): 931–945. Bibcode:2004Natur.431..931H. doi:10.1038/nature03001. PMID 15496913.
- ^ Braslavsky, Ido; et al. (2003). "Sequence information can be obtained from single DNA molecules". Proc Natl Acad Sci USA. 100 (7): 3960–3984. Bibcode:2003PNAS..100.3960B. doi:10.1073/pnas.0230489100. PMC 153030. PMID 12651960.
- ^ Heger, Monica (October 2, 2013). "Single-cell Sequencing Makes Strides in the Clinic with Cancer and PGD First Applications". Clinical Sequencing News.
- ^ Yurkiewicz, I. R.; Korf, B. R.; Lehmann, L. S. (2014). "Prenatal whole-genome sequencing--is the quest to know a fetus's future ethical?". New England Journal of Medicine. 370 (3): 195–7. doi:10.1056/NEJMp1215536. PMID 24428465.
- ^ Staden R (June 1979). "A strategy of DNA sequencing employing computer programs". Nucleic Acids Res. 6 (7): 2601–10. doi:10.1093/nar/6.7.2601. PMC 327874. PMID 461197.
- ^ Edwards, A; Caskey, T (1991). "Closure strategies for random DNA sequencing". Methods: A Companion to Methods in Enzymology. 3 (1): 41–47. doi:10.1016/S1046-2023(05)80162-8.
- ^ Edwards A; Voss H; Rice P; Civitello A; Stegemann J; Schwager C; Zimmermann J; Erfle H; Caskey CT; Ansorge W (April 1990). "Automated DNA sequencing of the human HPRT locus". Genomics. 6 (4): 593–608. doi:10.1016/0888-7543(90)90493-E. PMID 2341149.
- ^ Roach JC; Boysen C; Wang K; Hood L (March 1995). "Pairwise end sequencing: a unified approach to genomic mapping and sequencing". Genomics. 26 (2): 345–53. doi:10.1016/0888-7543(95)80219-C. PMID 7601461.
- ^ Fleischmann RD; Adams MD; White O; Clayton RA; Kirkness EF; Kerlavage AR; Bult CJ; Tomb JF; Dougherty BA; Merrick JM; McKenney; Sutton; Fitzhugh; Fields; Gocyne; Scott; Shirley; Liu; Glodek; Kelley; Weidman; Phillips; Spriggs; Hedblom; Cotton; Utterback; Hanna; Nguyen; Saudek; et al. (July 1995). "Whole-genome random sequencing and assembly of Haemophilus influenzae Rd". Science. 269 (5223): 496–512. Bibcode:1995Sci...269..496F. doi:10.1126/science.7542800. PMID 7542800.
- ^ Adams, MD; et al. (2000). "The genome sequence of Drosophila melanogaster". Science. 287 (5461): 2185–95. Bibcode:2000Sci...287.2185.. CiteSeerX 10.1.1.549.8639. doi:10.1126/science.287.5461.2185. PMID 10731132.
- ^ أ ب Mukhopadhyay R (February 2009). "DNA sequencers: the next generation". Anal. Chem. 81 (5): 1736–40. doi:10.1021/ac802712u. PMID 19193124.
- ^ Kwong, JC; McCallum, N; Sintchenko, V; Howden, BP (April 2015). "Whole genome sequencing in clinical and public health microbiology". Pathology. 47 (3): 199–210. doi:10.1097/pat.0000000000000235. PMC 4389090. PMID 25730631.
- ^ Strickland, Eliza (2015-10-14). "New Genetic Technologies Diagnose Critically Ill Infants Within 26 Hours - IEEE Spectrum". Spectrum.ieee.org. Archived from the original on 2015-11-16. Retrieved 2016-11-11.
- ^ "Article : Race to Cut Whole Genome Sequencing Costs Genetic Engineering & Biotechnology News — Biotechnology from Bench to Business". Genengnews.com. Archived from the original on 2006-10-17. Retrieved 2009-02-23.
- ^ "Whole Genome Sequencing Costs Continue to Drop". Eyeondna.com. Archived from the original on 2009-03-25. Retrieved 2009-02-23.
- ^ Harmon, Katherine (2010-06-28). "Genome Sequencing for the Rest of Us". Scientific American. Archived from the original on 2011-03-19. Retrieved 2010-08-13.
- ^ San Diego/Orange County Technology News. "Sequenom to Develop Third-Generation Nanopore-Based Single Molecule Sequencing Technology". Freshnews.com. Archived from the original on 2008-12-05. Retrieved 2009-02-24.
- ^ أ ب "Article : Whole Genome Sequencing in 24 Hours Genetic Engineering & Biotechnology News — Biotechnology from Bench to Business". Genengnews.com. Archived from the original on 2006-10-17. Retrieved 2009-02-23.
- ^ "Pacific Bio lifts the veil on its high-speed genome-sequencing effort". VentureBeat. 2008-02-10. Archived from the original on 2009-02-20. Retrieved 2009-02-23.
- ^ "Bio-IT World". Bio-IT World. 2008-10-06. Archived from the original on 2009-02-17. Retrieved 2009-02-23.
- ^ "With New Machine, Helicos Brings Personal Genome Sequencing A Step Closer". Xconomy. 2008-04-22. Archived from the original on 2011-01-02. Retrieved 2011-01-28.
- ^ "Whole genome sequencing costs continue to fall: $300 million in 2003, $1 million 2007, $60,000 now, $5000 by year end". Nextbigfuture.com. 2008-03-25. Archived from the original on 2010-12-20. Retrieved 2011-01-28.
- ^ "Han Cao's nanofluidic chip could cut DNA sequencing costs dramatically". Technology Review. Archived from the original on 2011-03-29.
- ^ Julia Karow (26 October 2015). "BGI Launches Desktop Sequencer in China; Plans to Register Platform With CFDA". GenomeWeb. Retrieved 2 December 2018.
- ^ "BGI Launches New Desktop Sequencer in China, Registers Larger Version With CFDA". 360Dx. GenomeWeb. 11 November 2016. Retrieved 2 December 2018.
- ^ Monica Heger (26 October 2018). "BGI Launches New Sequencer as Customers Report Data From Earlier Instruments". GenomeWeb. Retrieved 2 December 2018.
- ^ John Carroll (2008-07-14). "Pacific Biosciences gains $100M for sequencing tech". FierceBiotech. Archived from the original on 2009-05-01. Retrieved 2009-02-23.
- ^ Sibley, Lisa (2009-02-08). "Complete Genomics brings radical reduction in cost". Silicon Valley / San Jose Business Journal. Retrieved 2009-02-23.
- ^ Sarah Neville (5 March 2018). "Cheaper DNA sequencing unlocks secrets of rare diseases". Financial Times. Retrieved 2 December 2018.
- ^ Carlson, Rob (2007-01-02). "A Few Thoughts on Rapid Genome Sequencing and The Archon Prize — synthesis". Synthesis.cc. Archived from the original on 2009-08-08. Retrieved 2009-02-23.
- ^ "PRIZE Overview: Archon X PRIZE for Genomics".
- ^ Diamandis, Peter. "Outpaced by Innovation: Canceling an XPRIZE". Huffington Post. Archived from the original on 2013-08-25.
- ^ Aldhous, Peter. "X Prize for genomes cancelled before it begins". Archived from the original on 2016-09-21.
- ^ "SOLiD System — a next-gen DNA sequencing platform announced". Gizmag.com. 2007-10-27. Archived from the original on 2008-07-19. Retrieved 2009-02-24.
- ^ "The $1000 Genome: Coming Soon?". Dddmag.com. 2010-04-01. Archived from the original on 2011-04-15. Retrieved 2011-01-28.
- ^ "Individual genome sequencing — Illumina, Inc". Everygenome.com. Archived from the original on 2011-10-19. Retrieved 2011-01-28.
- ^ "Illumina launches personal genome sequencing service for $48,000 : Genetic Future". Scienceblogs.com. Archived from the original on June 16, 2009. Retrieved 2011-01-28.
- ^ Wade, Nicholas (2009-08-11). "Cost of Decoding a Genome Is Lowered". The New York Times. Archived from the original on 2013-05-21. Retrieved 2010-05-03.
- ^ "Technology Index". ABC News. Archived from the original on 15 May 2016. Retrieved 29 April 2018.
{{cite web}}: CS1 maint: bot: original URL status unknown (link) - ^ Drmanac R, Sparks AB, Callow MJ, et al. (2010). "Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays". Science. 327 (5961): 78–81. Bibcode:2010Sci...327...78D. doi:10.1126/science.1181498. PMID 19892942. S2CID 17309571.
- ^ "Illumina Announces $5,000 Genome Pricing – Bio-IT World". Archived from the original on 2011-05-17.
- ^ "NHGRI Awards More than $50M for Low-Cost DNA Sequencing Tech Development". Genome Web. 2009. Archived from the original on 2011-07-03.
- ^ "Life Technologies Introduces the Benchtop Ion Proton™ Sequencer; Designed to Decode a Human Genome in One Day for $1,000" (press release). Archived from the original on December 23, 2012. Retrieved August 30, 2012.
- ^ ANDREW POLLACK (2012-02-17). "Oxford Nanopore Unveils Tiny DNA Sequencing Device – The New York Times". The New York Times. Archived from the original on 2013-01-07. Retrieved 2016-11-11.
- ^ "Illumina Sequencer Enables $1,000 Genome". News: Genomics & Proteomics. Gen. Eng. Biotechnol. News (paper). Vol. 34, no. 4. 15 February 2014. p. 18.
- ^ Check Hayden, Erika (15 January 2014). "Is the $1,000 genome for real?". Nature: nature.2014.14530. doi:10.1038/nature.2014.14530. S2CID 211730238.
- ^ "The Cost of Sequencing a Human Genome". www.genome.gov. Archived from the original on 2016-11-25.
- ^ "With $999 Whole-Genome Sequencing Service, Veritas Embarks on Goal to Democratize DNA Information". 6 March 2016.
- ^ Andrews, Joe (2019-07-01). "23andMe competitor Veritas Genetics slashes price of whole genome sequencing 40% to $600". CNBC (in الإنجليزية). Retrieved 2019-09-02.
- ^ Megan Molteni (18 May 2017). "A Chinese Genome Giant Sets Its Sights on the Ultimate Sequencer". Wired. Retrieved 2 December 2018.
- ^ Phillips, K. A; Pletcher, M. J; Ladabaum, U (2015). "Is the "$1000 Genome" Really $1000? Understanding the Full Benefits and Costs of Genomic Sequencing". Technology and Health Care. 23 (3): 373–379. doi:10.3233/THC-150900. PMC 4527943. PMID 25669213.
- ^ "Blog: True Size of a Human Genome | Veritas Genetics". 28 July 2017.
- ^ "Psst, the human genome was never completely sequenced". statnews.com. 2017-06-20.
- ^ "Genomics Core". Gladstone.ucsf.edu. Archived from the original on June 30, 2010. Retrieved 2009-02-23.
- ^ Nishida N; Koike A; Tajima A; Ogasawara Y; Ishibashi Y; Uehara Y; Inoue I; Tokunaga K (2008). "Evaluating the performance of Affymetrix SNP Array 6.0 platform with 400 Japanese individuals". BMC Genomics. 9 (1): 431. doi:10.1186/1471-2164-9-431. PMC 2566316. PMID 18803882.
- ^ Petrone, Justin (16 January 2007). "Illumina, DeCode Build 1M SNP Chip; Q2 Launch to Coincide with Release of Affy's 6.0 SNP Array | BioArray News | Arrays". GenomeWeb. Archived from the original on 2011-07-16. Retrieved 2009-02-23.
- ^ Roach JC; Glusman G; Smit AF; et al. (April 2010). "Analysis of genetic inheritance in a family quartet by whole-genome sequencing". Science. 328 (5978): 636–9. Bibcode:2010Sci...328..636R. doi:10.1126/science.1186802. PMC 3037280. PMID 20220176.
- ^ Campbell CD; Chong JX; Malig M; et al. (November 2012). "Estimating the human mutation rate using autozygosity in a founder population". Nat. Genet. 44 (11): 1277–81. doi:10.1038/ng.2418. PMC 3483378. PMID 23001126.
- ^ Ye K; Beekman M; Lameijer EW; Zhang Y; Moed MH; van den Akker EB; Deelen J; Houwing-Duistermaat JJ; Kremer D; Anvar SY; Laros JF; Jones D; Raine K; Blackburne B; Potluri S; Long Q; Guryev V; van der Breggen R; Westendorp RG; 't Hoen PA; den Dunnen J; van Ommen GJ; Willemsen G; Pitts SJ; Cox DR; Ning Z; Boomsma DI; Slagboom PE (December 2013). "Aging as accelerated accumulation of somatic variants: whole-genome sequencing of centenarian and middle-aged monozygotic twin pairs". Twin Res Hum Genet. 16 (6): 1026–32. doi:10.1017/thg.2013.73. PMID 24182360.
- ^ Keightley PD (February 2012). "Rates and fitness consequences of new mutations in humans". Genetics. 190 (2): 295–304. doi:10.1534/genetics.111.134668. PMC 3276617. PMID 22345605.
- ^ Tuna M; Amos CI (November 2013). "Genomic sequencing in cancer". Cancer Lett. 340 (2): 161–70. doi:10.1016/j.canlet.2012.11.004. PMC 3622788. PMID 23178448.
- ^ Moran, Laurence A. (24 March 2011). "Sandwalk: How Big Is the Human Genome?". sandwalk.blogspot.com. Archived from the original on 1 December 2017. Retrieved 29 April 2018.
- ^ Hodgkinson, Alan; Chen, Ying; Eyre-Walker, Adam (January 2012). "The large-scale distribution of somatic mutations in cancer genomes". Human Mutation. 33 (1): 136–143. doi:10.1002/humu.21616. ISSN 1098-1004. PMID 21953857.
- ^ Supek, Fran; Lehner, Ben (2015-05-07). "Differential DNA mismatch repair underlies mutation rate variation across the human genome". Nature. 521 (7550): 81–84. Bibcode:2015Natur.521...81S. doi:10.1038/nature14173. ISSN 0028-0836. PMC 4425546. PMID 25707793.
- ^ Schuster-Böckler, Benjamin; Lehner, Ben (2012-08-23). "Chromatin organization is a major influence on regional mutation rates in human cancer cells". Nature. 488 (7412): 504–507. Bibcode:2012Natur.488..504S. doi:10.1038/nature11273. ISSN 1476-4687. PMID 22820252.
- ^ Supek, Fran; Lehner, Ben (2017-07-27). "Clustered Mutation Signatures Reveal that Error-Prone DNA Repair Targets Mutations to Active Genes". Cell. 170 (3): 534–547.e23. doi:10.1016/j.cell.2017.07.003. hdl:10230/35343. ISSN 1097-4172. PMID 28753428.
- ^ Yano, K; Yamamoto, E; Aya, K; Takeuchi, H; Lo, PC; Hu, L; Yamasaki, M; Yoshida, S; Kitano, H; Hirano, K; Matsuoka, M (August 2016). "Genome-wide association study using whole-genome sequencing rapidly identifies new genes influencing agronomic traits in rice". Nature Genetics. 48 (8): 927–34. doi:10.1038/ng.3596. PMID 27322545.
- ^ أ ب ت Abbott, Phil (2010). "US clinics quietly embrace whole-genome sequencing : Nature News". Nature. doi:10.1038/news.2010.465. Archived from the original on 2017-04-16. Retrieved 2016-11-11.
- ^ "One In A Billion: A boy's life, a medical mystery". Jsonline.com. Archived from the original on 2013-10-05. Retrieved 2016-11-11.
- ^ Mayer AN, Dimmock DP, Arca MJ, et al. (March 2011). "A timely arrival for genomic medicine". Genet. Med. 13 (3): 195–6. doi:10.1097/GIM.0b013e3182095089. PMID 21169843.
- ^ "Introducing diagnostic applications of '3Gb-testing' in human genetics". Archived from the original on 2014-11-10.
- ^ Boccia S, Mc Kee M, Adany R, Boffetta P, Burton H, Cambon-Thomsen A, Cornel MC, Gray M, Jani A, Knoppers BM, Khoury MJ, Meslin EM, Van Duijn CM, Villari P, Zimmern R, Cesario A, Puggina A, Colotto M, Ricciardi W (Aug 2014). "Beyond public health genomics: proposals from an international working group". Eur J Public Health. 24 (6): 877–879. doi:10.1093/eurpub/cku142. PMC 4245010. PMID 25168910.
- ^ "RD-Connect News 18 July 2014". Rd-connect.eu. Archived from the original on 10 October 2016. Retrieved 2016-11-11.
- ^ "Genomes2People: A Roadmap for Genomic Medicine". www.frontlinegenomics.com. Archived from the original on 14 February 2017. Retrieved 29 April 2018.
- ^ "The Risk Evaluation and Education for Alzheimer's Disease (REVEAL) Study - HBHE Genetics Research Group". hbhegenetics.sph.umich.edu. Archived from the original on 29 September 2017. Retrieved 29 April 2018.
- ^ "Risk Evaluation and Education for Alzheimer's Disease (REVEAL) II - Full Text View - ClinicalTrials.gov". clinicaltrials.gov. Archived from the original on 14 February 2017. Retrieved 29 April 2018.
- ^ Farnaes L; Hildreth A; Sweeney NM; Clark MM; Chowdhury S; Nahas S; Cakici JA; Benson W; Kaplan RH; Kronick R; Bainbridge MN; Friedman J; Gold JJ; Ding Y; Veeraraghavan N; Dimmock D; Kingsmore SF (2018). "Rapid whole-genome sequencing decreases infant morbidity and cost of hospitalization". NPJ Genomic Medicine. 3: 10. doi:10.1038/s41525-018-0049-4. PMC 5884823. PMID 29644095.
- ^ Wade, Nicholas (September 4, 2007). "In the Genome Race, the Sequel Is Personal". The New York Times. Archived from the original on April 11, 2009. Retrieved February 22, 2009.
- ^ Ledford, Heidi (2007). "Access : All about Craig: the first 'full' genome sequence". Nature. 449 (7158): 6–7. Bibcode:2007Natur.449....6L. doi:10.1038/449006a. PMID 17805257.
- ^ Levy S, Sutton G, Ng PC, Feuk L, Halpern AL, Walenz BP, Axelrod N, Huang J, Kirkness EF, Denisov G, Lin Y, MacDonald JR, Pang AW, Shago M, Stockwell TB, Tsiamouri A, Bafna V, Bansal V, Kravitz SA, Busam DA, Beeson KY, McIntosh TC, Remington KA, Abril JF, Gill J, Borman J, Rogers YH, Frazier ME, Scherer SW, Strausberg RL, Venter JC (September 2007). "The diploid genome sequence of an individual human". PLOS Biol. 5 (10): e254. doi:10.1371/journal.pbio.0050254. PMC 1964779. PMID 17803354.
- ^ Wade, Wade (June 1, 2007). "DNA pioneer Watson gets own genome map". International Herald Tribune. Archived from the original on September 27, 2008. Retrieved February 22, 2009.
- ^ Wade, Nicholas (May 31, 2007). "Genome of DNA Pioneer Is Deciphered". The New York Times. Archived from the original on June 20, 2011. Retrieved February 21, 2009.
- ^ Wheeler DA; Srinivasan M; Egholm M; Shen Y; Chen L; McGuire A; He W; Chen YJ; Makhijani V; Roth GT; Gomes X; Tartaro K; Niazi F; Turcotte CL; Irzyk GP; Lupski JR; Chinault C; Song XZ; Liu Y; Yuan Y; Nazareth L; Qin X; Muzny DM; Margulies M; Weinstock GM; Gibbs RA; Rothberg JM (2008). "The complete genome of an individual by massively parallel DNA sequencing". Nature. 452 (7189): 872–6. Bibcode:2008Natur.452..872W. doi:10.1038/nature06884. PMID 18421352.
- ^ Wang J; Wang, Wei; Li, Ruiqiang; Li, Yingrui; Tian, Geng; Goodman, Laurie; Fan, Wei; Zhang, Junqing; Li, Jun; Zhang, Juanbin, Juanbin; Guo, Yiran, Yiran; Feng, Binxiao, Binxiao; Li, Heng, Heng; Lu, Yao, Yao; Fang, Xiaodong, Xiaodong; Liang, Huiqing, Huiqing; Du, Zhenglin, Zhenglin; Li, Dong, Dong; Zhao, Yiqing, Yiqing; Hu, Yujie, Yujie; Yang, Zhenzhen, Zhenzhen; Zheng, Hancheng, Hancheng; Hellmann, Ines, Ines; Inouye, Michael, Michael; Pool, John, John; Yi, Xin, Xin; Zhao, Jing, Jing; Duan, Jinjie, Jinjie; Zhou, Yan, Yan; et al. (2008). "The diploid genome sequence of an Asian individual". Nature. 456 (7218): 60–65. Bibcode:2008Natur.456...60W. doi:10.1038/nature07484. PMC 2716080. PMID 18987735.
- ^ Bentley DR; Balasubramanian S; et al. (2008). "Accurate whole human genome sequencing using reversible terminator chemistry". Nature. 456 (7218): 53–9. Bibcode:2008Natur.456...53B. doi:10.1038/nature07517. PMC 2581791. PMID 18987734.
- ^ Ley TJ; Mardis ER; Ding L; Fulton B; McLellan MD; Chen K; Dooling D; Dunford-Shore BH; McGrath S; Hickenbotham M; Cook L; Abbott R; Larson DE; Koboldt DC; Pohl C; Smith S; Hawkins A; Abbott S; Locke D; Hillier LW; Miner T; Fulton L; Magrini V; Wylie T; Glasscock J; Conyers J; Sander N; Shi X; Osborne JR; et al. (2008). "DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome". Nature. 456 (7218): 66–72. Bibcode:2008Natur.456...66L. doi:10.1038/nature07485. PMC 2603574. PMID 18987736.
- ^ Lohr, Steve (2011-10-20). "New Book Details Jobs's Fight Against Cancer". The New York Times. Archived from the original on 2017-09-28.
- ^ "Complete Human Genome Sequencing Datasets to its Public Genomic Repository". Archived from the original on June 10, 2012.
- ^ Corpas M, Cariaso M, Coletta A, Weiss D, Harrison AP, Moran F, Yang H (November 12, 2013). "A Complete Public Domain Family Genomics Dataset". bioRxiv 10.1101/000216.
- ^ أ ب "200,000 whole genomes made available for biomedical studies by U.K. effort". www.science.org (in الإنجليزية). Retrieved 11 December 2021.
- ^ "Whole Genome Sequencing data on 200,000 UK Biobank participants available now". www.ukbiobank.ac.uk. 17 November 2021. Retrieved 11 December 2021.
- ^ Callaway, Ewen (2023-11-30). "World's biggest set of human genome sequences opens to scientists". Nature (in الإنجليزية). 624 (7990): 16–17. doi:10.1038/d41586-023-03763-3. PMID 38036674.
وصلات خارجية
- CS1 الإنجليزية الأمريكية-language sources (en-us)
- CS1 maint: bot: original URL status unknown
- Missing redirects
- Articles with hatnote templates targeting a nonexistent page
- Articles needing additional references from February 2017
- All articles needing additional references
- مقالات فيها عبارات متقادمة منذ يونيو 2012
- جميع المقالات التي فيها عبارات متقادمة
- تقانة حيوية
- علم الجينوم
- علم الأحياء الجزيئي
- معلوماتية حيوية
- دنا
- علم الوراثة الطبي
- Gene tests
- تقنيات ناشئة
- علم الوراثة الجزيئي